EnsembleVoteClassifier: A majority voting classifier
Implementation of a majority voting EnsembleVoteClassifier for classification.
from mlxtend.classifier import EnsembleVoteClassifier
Overview
The EnsembleVoteClassifier is a meta-classifier for combining similar or conceptually different machine learning classifiers for classification via majority or plurality voting. (For simplicity, we will refer to both majority and plurality voting as majority voting.)
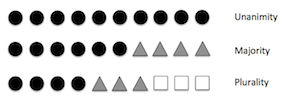

The EnsembleVoteClassifier implements "hard" and "soft" voting. In hard voting, we predict the final class label as the class label that has been predicted most frequently by the classification models. In soft voting, we predict the class labels by averaging the class-probabilities (only recommended if the classifiers are well-calibrated).

Note
If you are interested in using the EnsembleVoteClassifier, please note that it is now also available through scikit learn (>0.17) as VotingClassifier.
Majority Voting / Hard Voting
Hard voting is the simplest case of majority voting. Here, we predict the class label via majority (plurality) voting of each classifier :
Assuming that we combine three classifiers that classify a training sample as follows:
- classifier 1 -> class 0
- classifier 2 -> class 0
- classifier 3 -> class 1
Via majority vote, we would classify the sample as "class 0."
Weighted Majority Vote
In addition to the simple majority vote (hard voting) as described in the previous section, we can compute a weighted majority vote by associating a weight with classifier :
where is the characteristic function , and is the set of unique class labels.
Continuing with the example from the previous section
- classifier 1 -> class 0
- classifier 2 -> class 0
- classifier 3 -> class 1
assigning the weights {0.2, 0.2, 0.6} would yield a prediction :
Soft Voting
In soft voting, we predict the class labels based on the predicted probabilities for classifier -- this approach is only recommended if the classifiers are well-calibrated.
where is the weight that can be assigned to the th classifier.
Assuming the example in the previous section was a binary classification task with class labels , our ensemble could make the following prediction:
Using uniform weights, we compute the average probabilities:
However, assigning the weights {0.1, 0.1, 0.8} would yield a prediction :
References
- [1] S. Raschka. Python Machine Learning. Packt Publishing Ltd., 2015.
Example 1 - Classifying Iris Flowers Using Different Classification Models
from sklearn import datasets
iris = datasets.load_iris()
X, y = iris.data[:, 1:3], iris.target
from sklearn import model_selection
from sklearn.linear_model import LogisticRegression
from sklearn.naive_bayes import GaussianNB
from sklearn.ensemble import RandomForestClassifier
import numpy as np
clf1 = LogisticRegression(random_state=1)
clf2 = RandomForestClassifier(random_state=1)
clf3 = GaussianNB()
print('5-fold cross validation:\n')
labels = ['Logistic Regression', 'Random Forest', 'Naive Bayes']
for clf, label in zip([clf1, clf2, clf3], labels):
scores = model_selection.cross_val_score(clf, X, y,
cv=5,
scoring='accuracy')
print("Accuracy: %0.2f (+/- %0.2f) [%s]"
% (scores.mean(), scores.std(), label))
5-fold cross validation:
Accuracy: 0.95 (+/- 0.04) [Logistic Regression]
Accuracy: 0.94 (+/- 0.04) [Random Forest]
Accuracy: 0.91 (+/- 0.04) [Naive Bayes]
from mlxtend.classifier import EnsembleVoteClassifier
eclf = EnsembleVoteClassifier(clfs=[clf1, clf2, clf3], weights=[1,1,1])
labels = ['Logistic Regression', 'Random Forest', 'Naive Bayes', 'Ensemble']
for clf, label in zip([clf1, clf2, clf3, eclf], labels):
scores = model_selection.cross_val_score(clf, X, y,
cv=5,
scoring='accuracy')
print("Accuracy: %0.2f (+/- %0.2f) [%s]"
% (scores.mean(), scores.std(), label))
Accuracy: 0.95 (+/- 0.04) [Logistic Regression]
Accuracy: 0.94 (+/- 0.04) [Random Forest]
Accuracy: 0.91 (+/- 0.04) [Naive Bayes]
Accuracy: 0.95 (+/- 0.04) [Ensemble]
Plotting Decision Regions
import matplotlib.pyplot as plt
from mlxtend.plotting import plot_decision_regions
import matplotlib.gridspec as gridspec
import itertools
gs = gridspec.GridSpec(2, 2)
fig = plt.figure(figsize=(10,8))
labels = ['Logistic Regression', 'Random Forest', 'Naive Bayes', 'Ensemble']
for clf, lab, grd in zip([clf1, clf2, clf3, eclf],
labels,
itertools.product([0, 1], repeat=2)):
clf.fit(X, y)
ax = plt.subplot(gs[grd[0], grd[1]])
fig = plot_decision_regions(X=X, y=y, clf=clf)
plt.title(lab)
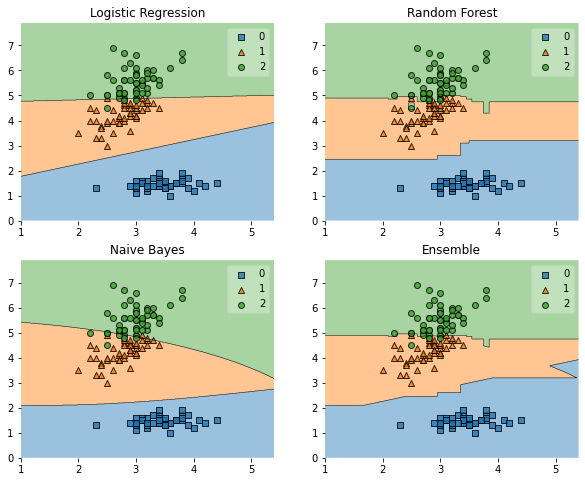
Example 2 - Grid Search
from sklearn import datasets
iris = datasets.load_iris()
X, y = iris.data[:, 1:3], iris.target
%%capture --no-display
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import LogisticRegression
from sklearn.naive_bayes import GaussianNB
from sklearn.ensemble import RandomForestClassifier
from mlxtend.classifier import EnsembleVoteClassifier
clf1 = LogisticRegression(random_state=1)
clf2 = RandomForestClassifier(random_state=1)
clf3 = GaussianNB()
eclf = EnsembleVoteClassifier(clfs=[clf1, clf2, clf3], voting='soft')
params = {'logisticregression__C': [1.0, 100.0],
'randomforestclassifier__n_estimators': [20, 200],}
grid = GridSearchCV(estimator=eclf, param_grid=params, cv=5)
grid.fit(iris.data, iris.target)
cv_keys = ('mean_test_score', 'std_test_score', 'params')
for r, _ in enumerate(grid.cv_results_['mean_test_score']):
print("%0.3f +/- %0.2f %r"
% (grid.cv_results_[cv_keys[0]][r],
grid.cv_results_[cv_keys[1]][r] / 2.0,
grid.cv_results_[cv_keys[2]][r]))
Note: If the EnsembleClassifier is initialized with multiple similar estimator objects, the estimator names are modified with consecutive integer indices, for example:
%%capture --no-display
clf1 = LogisticRegression(random_state=1)
clf2 = RandomForestClassifier(random_state=1)
eclf = EnsembleVoteClassifier(clfs=[clf1, clf1, clf2],
voting='soft')
params = {'logisticregression-1__C': [1.0, 100.0],
'logisticregression-2__C': [1.0, 100.0],
'randomforestclassifier__n_estimators': [20, 200],}
grid = GridSearchCV(estimator=eclf, param_grid=params, cv=5)
grid = grid.fit(iris.data, iris.target)
Note
The EnsembleVoteClass also enables grid search over the clfs argument. However, due to the current implementation of GridSearchCV in scikit-learn, it is not possible to search over both, differenct classifiers and classifier parameters at the same time. For instance, while the following parameter dictionary works
params = {'randomforestclassifier__n_estimators': [1, 100],
'clfs': [(clf1, clf1, clf1), (clf2, clf3)]}
it will use the instance settings of clf1, clf2, and clf3 and not overwrite it with the 'n_estimators' settings from 'randomforestclassifier__n_estimators': [1, 100].
Example 3 - Majority voting with classifiers trained on different feature subsets
Feature selection algorithms implemented in scikit-learn as well as the SequentialFeatureSelector implement a transform method that passes the reduced feature subset to the next item in a Pipeline.
For example, the method
def transform(self, X):
return X[:, self.k_feature_idx_]
returns the best feature columns, k_feature_idx_, given a dataset X.
Thus, we simply need to construct a Pipeline consisting of the feature selector and the classifier in order to select different feature subsets for different algorithms. During fitting, the optimal feature subsets are automatically determined via the GridSearchCV object, and by calling predict, the fitted feature selector in the pipeline only passes these columns along, which resulted in the best performance for the respective classifier.
%%capture --no-display
from sklearn import datasets
iris = datasets.load_iris()
X, y = iris.data[:, :], iris.target
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import LogisticRegression
from sklearn.naive_bayes import GaussianNB
from sklearn.ensemble import RandomForestClassifier
from mlxtend.classifier import EnsembleVoteClassifier
from sklearn.pipeline import Pipeline
from mlxtend.feature_selection import SequentialFeatureSelector
clf1 = LogisticRegression(random_state=1)
clf2 = RandomForestClassifier(random_state=1)
clf3 = GaussianNB()
# Creating a feature-selection-classifier pipeline
sfs1 = SequentialFeatureSelector(clf1,
k_features=4,
forward=True,
floating=False,
scoring='accuracy',
verbose=0,
cv=0)
clf1_pipe = Pipeline([('sfs', sfs1),
('logreg', clf1)])
eclf = EnsembleVoteClassifier(clfs=[clf1_pipe, clf2, clf3],
voting='soft')
params = {'pipeline__sfs__k_features': [1, 2, 3],
'pipeline__logreg__C': [1.0, 100.0],
'randomforestclassifier__n_estimators': [20, 200]}
grid = GridSearchCV(estimator=eclf, param_grid=params, cv=5)
grid.fit(iris.data, iris.target)
cv_keys = ('mean_test_score', 'std_test_score', 'params')
for r, _ in enumerate(grid.cv_results_['mean_test_score']):
print("%0.3f +/- %0.2f %r"
% (grid.cv_results_[cv_keys[0]][r],
grid.cv_results_[cv_keys[1]][r] / 2.0,
grid.cv_results_[cv_keys[2]][r]))
The best parameters determined via GridSearch are:
grid.best_params_
{'pipeline__logreg__C': 1.0,
'pipeline__sfs__k_features': 2,
'randomforestclassifier__n_estimators': 200}
Now, we assign these parameters to the ensemble voting classifier, fit the models on the complete training set, and perform a prediction on 3 samples from the Iris dataset.
eclf = eclf.set_params(**grid.best_params_)
eclf.fit(X, y).predict(X[[1, 51, 149]])
array([0, 1, 2])
Manual Approach
Alternatively, we can select different columns "manually" using the ColumnSelector object. In this example, we select only the first (sepal length) and third (petal length) column for the logistic regression classifier (clf1).
from mlxtend.feature_selection import ColumnSelector
col_sel = ColumnSelector(cols=[0, 2])
clf1_pipe = Pipeline([('sel', col_sel),
('logreg', clf1)])
eclf = EnsembleVoteClassifier(clfs=[clf1_pipe, clf2, clf3],
voting='soft')
eclf.fit(X, y).predict(X[[1, 51, 149]])
array([0, 1, 2])
Furthermore, we can fit the SequentialFeatureSelector separately, outside the grid search hyperparameter optimization pipeline. Here, we determine the best features first, and then we construct a pipeline using these "fixed," best features as seed for the ColumnSelector:
sfs1 = SequentialFeatureSelector(clf1,
k_features=2,
forward=True,
floating=False,
scoring='accuracy',
verbose=1,
cv=0)
sfs1.fit(X, y)
print('Best features', sfs1.k_feature_idx_)
col_sel = ColumnSelector(cols=sfs1.k_feature_idx_)
clf1_pipe = Pipeline([('sel', col_sel),
('logreg', clf1)])
[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers.
[Parallel(n_jobs=1)]: Done 4 out of 4 | elapsed: 0.0s finished
Features: 1/2[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers.
[Parallel(n_jobs=1)]: Done 3 out of 3 | elapsed: 0.0s finished
Features: 2/2
Best features (2, 3)
eclf = EnsembleVoteClassifier(clfs=[clf1_pipe, clf2, clf3],
voting='soft')
eclf.fit(X, y).predict(X[[1, 51, 149]])
array([0, 1, 2])
Example 5 - Using Pre-fitted Classifiers
from sklearn import datasets
iris = datasets.load_iris()
X, y = iris.data[:, 1:3], iris.target
Assume that we previously fitted our classifiers:
from sklearn import model_selection
from sklearn.linear_model import LogisticRegression
from sklearn.naive_bayes import GaussianNB
from sklearn.ensemble import RandomForestClassifier
import numpy as np
clf1 = LogisticRegression(random_state=1)
clf2 = RandomForestClassifier(random_state=1)
clf3 = GaussianNB()
for clf in (clf1, clf2, clf3):
clf.fit(X, y)
By setting fit_base_estimators=False, it will enforce use_clones to be False and the EnsembleVoteClassifier will not re-fit these classifers to save computational time:
from mlxtend.classifier import EnsembleVoteClassifier
import copy
eclf = EnsembleVoteClassifier(clfs=[clf1, clf2, clf3], weights=[1,1,1], fit_base_estimators=False)
labels = ['Logistic Regression', 'Random Forest', 'Naive Bayes', 'Ensemble']
eclf.fit(X, y)
print('accuracy:', np.mean(y == eclf.predict(X)))
accuracy: 0.96
/Users/sebastian/miniforge3/lib/python3.9/site-packages/mlxtend/classifier/ensemble_vote.py:166: UserWarning: fit_base_estimators=False enforces use_clones to be `False`
warnings.warn("fit_base_estimators=False "
However, please note that fit_base_estimators=False is incompatible to any form of cross-validation that is done in e.g., model_selection.cross_val_score or model_selection.GridSearchCV, etc., since it would require the classifiers to be refit to the training folds. Thus, only use fit_base_estimators=False if you want to make a prediction directly without cross-validation.
Example 6 - Ensembles of Classifiers that Operate on Different Feature Subsets
If desired, the different classifiers can be fit to different subsets of features in the training dataset. The following example illustrates how this can be done on a technical level using scikit-learn pipelines and the ColumnSelector:
from sklearn.datasets import load_iris
from mlxtend.classifier import EnsembleVoteClassifier
from mlxtend.feature_selection import ColumnSelector
from sklearn.pipeline import make_pipeline
from sklearn.linear_model import LogisticRegression
iris = load_iris()
X = iris.data
y = iris.target
pipe1 = make_pipeline(ColumnSelector(cols=(0, 2)),
LogisticRegression())
pipe2 = make_pipeline(ColumnSelector(cols=(1, 2, 3)),
LogisticRegression())
eclf = EnsembleVoteClassifier(clfs=[pipe1, pipe2])
eclf.fit(X, y)
EnsembleVoteClassifier(clfs=[Pipeline(steps=[('columnselector',
ColumnSelector(cols=(0, 2))),
('logisticregression',
LogisticRegression())]),
Pipeline(steps=[('columnselector',
ColumnSelector(cols=(1, 2, 3))),
('logisticregression',
LogisticRegression())])])
Example 7 - A Note about Scikit-Learn SVMs and Soft Voting
This section provides some additional technical insights in how probabilities are used when voting='soft'.
Note that scikit-learn estimates the probabilities for SVMs (more info here: https://scikit-learn.org/stable/modules/svm.html#scores-probabilities) in a way that these may not be consistent with the class labels that the SVM predicts. This is an extreme example, but let's say we have a dataset with 3 class labels, 0, 1, and 2. For a given training example, the SVM classifier may predict class 2. However, the class-membership probabilities may look as follows:
- class 0: 99%
- class 1: 0.5%
- class 2: 0.5%
A practical example of this scenario is shown below:
import numpy as np
from mlxtend.classifier import EnsembleVoteClassifier
from sklearn.svm import SVC
from sklearn.datasets import load_iris
iris = load_iris()
X, y = iris.data, iris.target
clf2 = SVC(probability=True, random_state=4)
clf2.fit(X, y)
eclf = EnsembleVoteClassifier(clfs=[clf2], voting='soft', fit_base_estimators=False)
eclf.fit(X, y)
for svm_class, e_class, svm_prob, e_prob, in zip(clf2.predict(X),
eclf.predict(X),
clf2.predict_proba(X),
eclf.predict_proba(X)):
if svm_class != e_class:
print('============')
print('Probas from SVM :', svm_prob)
print('Class from SVM :', svm_class)
print('Probas from SVM in Ensemble:', e_prob)
print('Class from SVM in Ensemble :', e_class)
print('============')
============
Probas from SVM : [0.00921708 0.49415165 0.49663127]
Class from SVM : 1
Probas from SVM in Ensemble: [0.00921708 0.49415165 0.49663127]
Class from SVM in Ensemble : 2
============
/Users/sebastian/miniforge3/lib/python3.9/site-packages/mlxtend/classifier/ensemble_vote.py:166: UserWarning: fit_base_estimators=False enforces use_clones to be `False`
warnings.warn("fit_base_estimators=False "
Example 7 - A Note about Scikit-Learn SVMs and Soft Voting
Based on the probabilities, we would expect the SVM to predict class 2, because it has the highest probability. Since the EnsembleVoteClassifier uses the argmax function internally if voting='soft', it would indeed predict class 2 in this case even if the ensemble consists of only one SVM model.
Note that in practice, this minor technical detail does not need to concern you, but it is useful to keep it in mind in case you are wondering about results from a 1-model SVM ensemble compared to that SVM alone -- this is not a bug.
Example 8 - Optimizing Ensemble Weights with Nelder-Mead
In this section, we will see how we can use a heuristic search method like Nelder-Mead for optimizing the ensemble weights.
Suppose we have the following example scenario where we fit 3 individual classifiers on different subsets of the training dataset:
from mlxtend.classifier import EnsembleVoteClassifier
from mlxtend.data import mnist_data
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn.naive_bayes import GaussianNB
from sklearn.tree import DecisionTreeClassifier
X, y = mnist_data()
X_train, X_val, y_train, y_val = train_test_split(
X, y, test_size=0.5, shuffle=True, random_state=1
)
clf1 = GaussianNB()
clf2 = LogisticRegression(random_state=123, solver='newton-cg')
clf3 = DecisionTreeClassifier(random_state=123, max_depth=2)
clf1.fit(X_train[500:1000], y_train[500:1000])
clf2.fit(X_train[750:1250], y_train[750:1250])
clf3.fit(X_train[1250:2000], y_train[1250:2000]);
Then, we construct the an ensemble classifier from these 3 classifiers where each classifier contributes equally with weight 1:
eclf = EnsembleVoteClassifier(
clfs=(clf1, clf2, clf3),
voting="soft", # the same would also work with "hard" voting
weights=(1, 1, 1),
use_clones=False,
fit_base_estimators=False,
)
eclf.fit(X_train, y_train)
eclf.score(X_val, y_val)
/Users/sebastian/miniforge3/lib/python3.9/site-packages/mlxtend/classifier/ensemble_vote.py:166: UserWarning: fit_base_estimators=False enforces use_clones to be `False`
warnings.warn("fit_base_estimators=False "
0.8012
We see that we reach 80% accuracy on the validation set. Can we do better? Maybe they indvidual classifiers shouldn't be contributing equally. Perhaps, we can use an optimization algorothm from scipy.optimize to find a better relative weighting of these individual classifiers.
Let's set up an objective function that we want to minimize via SciPy's minimize:
from scipy.optimize import minimize
def function_to_minimize(weights, fitted_clfs):
w1, w2 = weights # these are the new weights!
newclf = EnsembleVoteClassifier(
voting="soft",
use_clones=False,
fit_base_estimators=False,
clfs=fitted_clfs,
weights=(w1, w2, 1.), # use the new weights
)
newclf.fit(X_train, y_train)
score = newclf.score(X_val, y_val)
# change accuracy to error so that smaller is better
score_to_minimize = 1 - score
return score_to_minimize
Note a few things:
-
We only optimize 2 out of the 3 classifier weigths. That's because the weighting is relative to each other, and it would be overkill (and too many degrees of freedom) to also optimize weight 3.
-
We set
use_clones=False & fit_base_estimators=Falseas before, this is to make sure that we use the prefit classifiers in the ensemble classifier. -
Instead of optimizing the accuracy, we optimize the the classification error,
score_to_minimize = 1 - score. That's because we use theminimizefunction where lower means better.
Next, let's choose some initial weight values and run the optimization. Via the bounds we specify the range (lower and upper value) for each weight so that the search doesn't go crazy:
%%capture --no-display
init_weights = [1., 1.]
results = minimize(
function_to_minimize,
init_weights,
args=((clf1, clf2, clf3),),
bounds=((0, 5), (0, 5)),
method="nelder-mead",
)
Let's look at the results!
print(results)
final_simplex: (array([[0.575 , 1.40625 ],
[0.57500153, 1.40622215],
[0.57508965, 1.40617647]]), array([0.1324, 0.1324, 0.1324]))
fun: 0.13239999999999996
message: 'Optimization terminated successfully.'
nfev: 60
nit: 21
status: 0
success: True
x: array([0.575 , 1.40625])
It looks like the search was successful and returned the following weights:
solution = results["x"]
print(solution)
[0.575 1.40625]
Let's use these new weights in our ensemble classifier:
eclf = EnsembleVoteClassifier(
clfs=(clf1, clf2, clf3),
voting="soft",
weights=(solution[0], solution[1], 1),
use_clones=False,
fit_base_estimators=False,
)
eclf.fit(X_train, y_train)
eclf.score(X_val, y_val)
/Users/sebastian/miniforge3/lib/python3.9/site-packages/mlxtend/classifier/ensemble_vote.py:166: UserWarning: fit_base_estimators=False enforces use_clones to be `False`
warnings.warn("fit_base_estimators=False "
0.8676
As we can see, the results on the validation set (0.8676) improved compared to the original ones (0.8012). Yay!
API
EnsembleVoteClassifier(clfs, voting='hard', weights=None, verbose=0, use_clones=True, fit_base_estimators=True)
Soft Voting/Majority Rule classifier for scikit-learn estimators.
Parameters
-
clfs: array-like, shape = [n_classifiers]A list of classifiers. Invoking the
fitmethod on theVotingClassifierwill fit clones of those original classifiers be stored in the class attribute ifuse_clones=True(default) andfit_base_estimators=True(default). -
voting: str, {'hard', 'soft'} (default='hard')If 'hard', uses predicted class labels for majority rule voting. Else if 'soft', predicts the class label based on the argmax of the sums of the predicted probalities, which is recommended for an ensemble of well-calibrated classifiers.
-
weights: array-like, shape = [n_classifiers], optional (default=None)Sequence of weights (
floatorint) to weight the occurances of predicted class labels (hardvoting) or class probabilities before averaging (softvoting). Uses uniform weights ifNone. -
verbose: int, optional (default=0)Controls the verbosity of the building process. -
verbose=0(default): Prints nothing -verbose=1: Prints the number & name of the clf being fitted -verbose=2: Prints info about the parameters of the clf being fitted -verbose>2: Changesverboseparam of the underlying clf to self.verbose - 2 -
use_clones: bool (default: True)Clones the classifiers for stacking classification if True (default) or else uses the original ones, which will be refitted on the dataset upon calling the
fitmethod. Hence, if use_clones=True, the original input classifiers will remain unmodified upon using the StackingClassifier'sfitmethod. Settinguse_clones=Falseis recommended if you are working with estimators that are supporting the scikit-learn fit/predict API interface but are not compatible to scikit-learn'sclonefunction. -
fit_base_estimators: bool (default: True)Refits classifiers in
clfsif True; uses references to theclfs, otherwise (assumes that the classifiers were already fit). Note: fit_base_estimators=False will enforce use_clones to be False, and is incompatible to most scikit-learn wrappers! For instance, if any form of cross-validation is performed this would require the re-fitting classifiers to training folds, which would raise a NotFitterError if fit_base_estimators=False. (New in mlxtend v0.6.)
Attributes
-
classes_: array-like, shape = [n_predictions] -
clf: array-like, shape = [n_predictions]The input classifiers; may be overwritten if
use_clones=False -
clf_: array-like, shape = [n_predictions]Fitted input classifiers; clones if
use_clones=True
Examples
```
>>> import numpy as np
>>> from sklearn.linear_model import LogisticRegression
>>> from sklearn.naive_bayes import GaussianNB
>>> from sklearn.ensemble import RandomForestClassifier
>>> from mlxtend.sklearn import EnsembleVoteClassifier
>>> clf1 = LogisticRegression(random_seed=1)
>>> clf2 = RandomForestClassifier(random_seed=1)
>>> clf3 = GaussianNB()
>>> X = np.array([[-1, -1], [-2, -1], [-3, -2], [1, 1], [2, 1], [3, 2]])
>>> y = np.array([1, 1, 1, 2, 2, 2])
>>> eclf1 = EnsembleVoteClassifier(clfs=[clf1, clf2, clf3],
... voting='hard', verbose=1)
>>> eclf1 = eclf1.fit(X, y)
>>> print(eclf1.predict(X))
[1 1 1 2 2 2]
>>> eclf2 = EnsembleVoteClassifier(clfs=[clf1, clf2, clf3], voting='soft')
>>> eclf2 = eclf2.fit(X, y)
>>> print(eclf2.predict(X))
[1 1 1 2 2 2]
>>> eclf3 = EnsembleVoteClassifier(clfs=[clf1, clf2, clf3],
... voting='soft', weights=[2,1,1])
>>> eclf3 = eclf3.fit(X, y)
>>> print(eclf3.predict(X))
[1 1 1 2 2 2]
>>>
For more usage examples, please see
https://rasbt.github.io/mlxtend/user_guide/classifier/EnsembleVoteClassifier/
```
Methods
fit(X, y, sample_weight=None)
Learn weight coefficients from training data for each classifier.
Parameters
-
X: {array-like, sparse matrix}, shape = [n_samples, n_features]Training vectors, where n_samples is the number of samples and n_features is the number of features.
-
y: array-like, shape = [n_samples]Target values.
-
sample_weight: array-like, shape = [n_samples], optionalSample weights passed as sample_weights to each regressor in the regressors list as well as the meta_regressor. Raises error if some regressor does not support sample_weight in the fit() method.
Returns
self: object
fit_transform(X, y=None, fit_params)
Fit to data, then transform it.
Fits transformer to `X` and `y` with optional parameters `fit_params`
and returns a transformed version of `X`.
Parameters
-
X: array-like of shape (n_samples, n_features)Input samples.
-
y: array-like of shape (n_samples,) or (n_samples, n_outputs), default=NoneTarget values (None for unsupervised transformations).
-
**fit_params: dictAdditional fit parameters.
Returns
-
X_new: ndarray array of shape (n_samples, n_features_new)Transformed array.
get_params(deep=True)
Return estimator parameter names for GridSearch support.
predict(X)
Predict class labels for X.
Parameters
-
X: {array-like, sparse matrix}, shape = [n_samples, n_features]Training vectors, where n_samples is the number of samples and n_features is the number of features.
Returns
-
maj: array-like, shape = [n_samples]Predicted class labels.
predict_proba(X)
Predict class probabilities for X.
Parameters
-
X: {array-like, sparse matrix}, shape = [n_samples, n_features]Training vectors, where n_samples is the number of samples and n_features is the number of features.
Returns
-
avg: array-like, shape = [n_samples, n_classes]Weighted average probability for each class per sample.
score(X, y, sample_weight=None)
Return the mean accuracy on the given test data and labels.
In multi-label classification, this is the subset accuracy
which is a harsh metric since you require for each sample that
each label set be correctly predicted.
Parameters
-
X: array-like of shape (n_samples, n_features)Test samples.
-
y: array-like of shape (n_samples,) or (n_samples, n_outputs)True labels for
X. -
sample_weight: array-like of shape (n_samples,), default=NoneSample weights.
Returns
-
score: floatMean accuracy of
self.predict(X)wrt.y.
set_params(params)
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects
(such as :class:`~sklearn.pipeline.Pipeline`). The latter have
parameters of the form ``<component>__<parameter>`` so that it's
possible to update each component of a nested object.
Parameters
-
**params: dictEstimator parameters.
Returns
-
self: estimator instanceEstimator instance.
transform(X)
Return class labels or probabilities for X for each estimator.
Parameters
-
X: {array-like, sparse matrix}, shape = [n_samples, n_features]Training vectors, where n_samples is the number of samples and n_features is the number of features.
Returns
-
Ifvoting='soft'`` : array-like = [n_classifiers, n_samples, n_classes]Class probabilties calculated by each classifier.
-
Ifvoting='hard'`` : array-like = [n_classifiers, n_samples]Class labels predicted by each classifier.
ython